Annotating Data With Metadata
Annotations help users search for and find data, and they are a powerful tool used to systematically group and/or describe things in Synapse.
Annotations are stored as key-value pairs in Synapse, where the key defines a particular aspect of your data (for example, species, assay, file format) and the value defines a variable that belongs to that category (mouse, RNAseq, .bam). You can use annotations to add additional information about a project, file, folder, table, or view. Annotations can be based on an existing ontology or controlled vocabulary, or can be created as needed and modified later as your metadata evolves.
For example, if you have uploaded a collection of alignment files in the BAM file format from an RNA-sequencing experiment, each representing a sample and experimental replicate, you can use annotations to surface this information in a structured way. Sometimes, users encode this information in file names, e.g., sampleA_conditionB.bam, which makes it “human-readable” but not searchable.
In this case, you may want to add annotations that look like this:
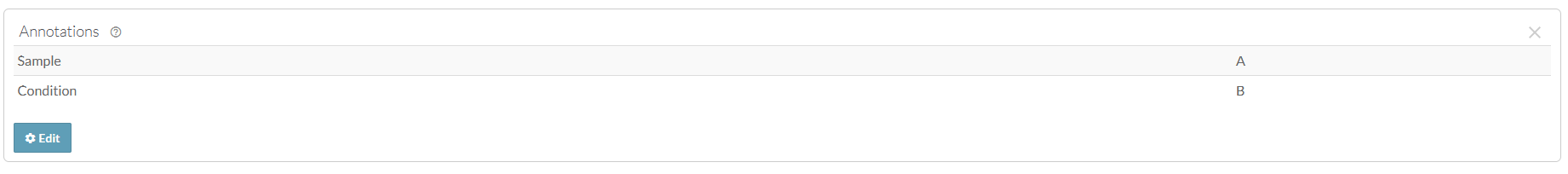
You can add and edit annotations from the web or programmatically using the command line client, the Python client, the R client. Using the programmatic clients facilitates batching and automated population of annotations across many files. The web client can be used to bulk update many files using views.Adding and Editing Annotations via the Synapse UI
To add or modify annotations on projects, files, folders, or tables in the web client, find the Tools menu in the upper right corner and select Annotations.
A new window will appear with a list of any previously added annotations. To add new annotations or edit existing annotations, click Edit.

In the pop-up window, add your annotations one at a time. Use the + icon to add multiple values for a single key and the x icon to remove values. Click Add New Key to add a new key.
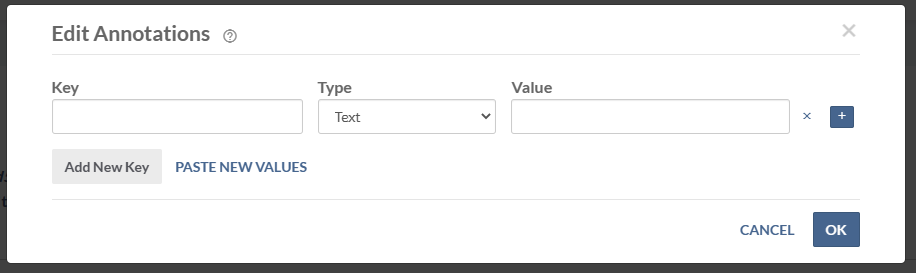
![]() To add annotations on multiple files, refer to Managing Custom Metadata at Scale for a tutorial on using views for annotation management.
To add annotations on multiple files, refer to Managing Custom Metadata at Scale for a tutorial on using views for annotation management.
Adding and Editing Annotations Programmatically
You can programmatically add annotations during file upload or after.
Command line
To add annotations on a new file during upload:
synapse store sampleA_conditionB.bam --parentId syn00123 --annotations '{"fileFormat":"bam", "assay":"rnaSeq"}'
To add annotations on an existing file:
synapse set-annotations --id syn00123 --annotations '{"fileFormat":"bam", "assay":"rnaSeq"}'
Python
To add annotations on a new file during upload:
entity = File(path="sampleA_conditionB.bam",parent="syn00123")
entity.annotations = {"fileFormat":"bam", "assay":"rnaSeq"}
syn.store(entity)
To modify annotations on an existing file:
entity = syn.get_annotations("syn123")
# set key 'fileFormat' to have value 'fastq'
entity['fileFormat'] = 'fastq'
syn.set_annotations(entity)
R
To add annotations on a new file during upload:
entity <- File("sampleA_conditionB.bam", parent="syn00123")
entity <- synStore(entity, annotations=list(fileFormat = "bam", assay = "rnaSeq"))
To modify annotations on an existing file:
entity <- synGet("syn00123")
##### Modify annotations and PRESERVE existing annotations
existing_annots <- synGetAnnotations(entity)
synSetAnnotations(entity, annotations = c(existing_annots, list(fileType = "bam", assay = "rnaSeq")))
##### Modify annotations and REMOVE existing annotations
synSetAnnotations(entity, annotations = list(fileType = "bam", assay = "rnaSeq"))
